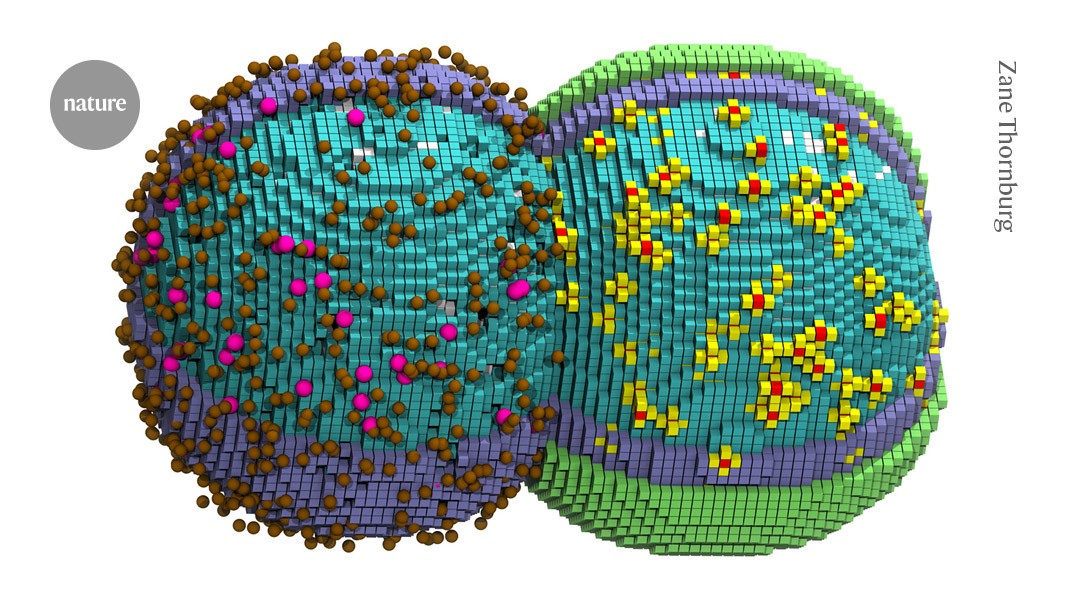
"To model bacterial life, Thornburg turned to one of its simplest examples: a bacterial cell with a 'minimal' genome. The organism, named JCVI-Syn3a, was created by whittling the genome of the parasite Mycoplasma mycoides down to just 493 genes, dispensing with more than 400 non-essential genes."
"Thornburg created a three-dimensional simulation that attempted to account for the cell's DNA, proteins, ribosomes, and other molecules of life, as they change over time. Specific molecules, such as a DNA-copying enzyme, obeyed rules based on real-world measurements, and reactions occurred when interacting partners became close in physical space."
"The team's goal was to simulate JCVI-Syn3a over the time it takes to copy its DNA and divide into two, known as the cell cycle. Some early attempts failed, Thornburg says, because the genome kept falling apart faster than it could be made, or spewing out of the cell membrane."
Researchers at the University of Illinois created the first comprehensive simulation of nearly all chemical reactions occurring within a living bacterial cell. Using JCVI-Syn3a, a bacterium with a minimal genome of only 493 genes, they built a three-dimensional model tracking DNA, proteins, ribosomes, and other molecular interactions over time. The simulation accounted for the complete cell cycle, from DNA copying through cell division. While some simplifications were necessary—such as modeling unknown gene functions as inert spheres and limiting ribosomes to one per mRNA transcript—the model successfully demonstrated how molecular interactions give rise to cellular life processes.
#cellular-simulation #bacterial-cell-modeling #molecular-biology #computational-biophysics #cell-cycle
Read at Nature
Unable to calculate read time
Collection
[
|
...
]